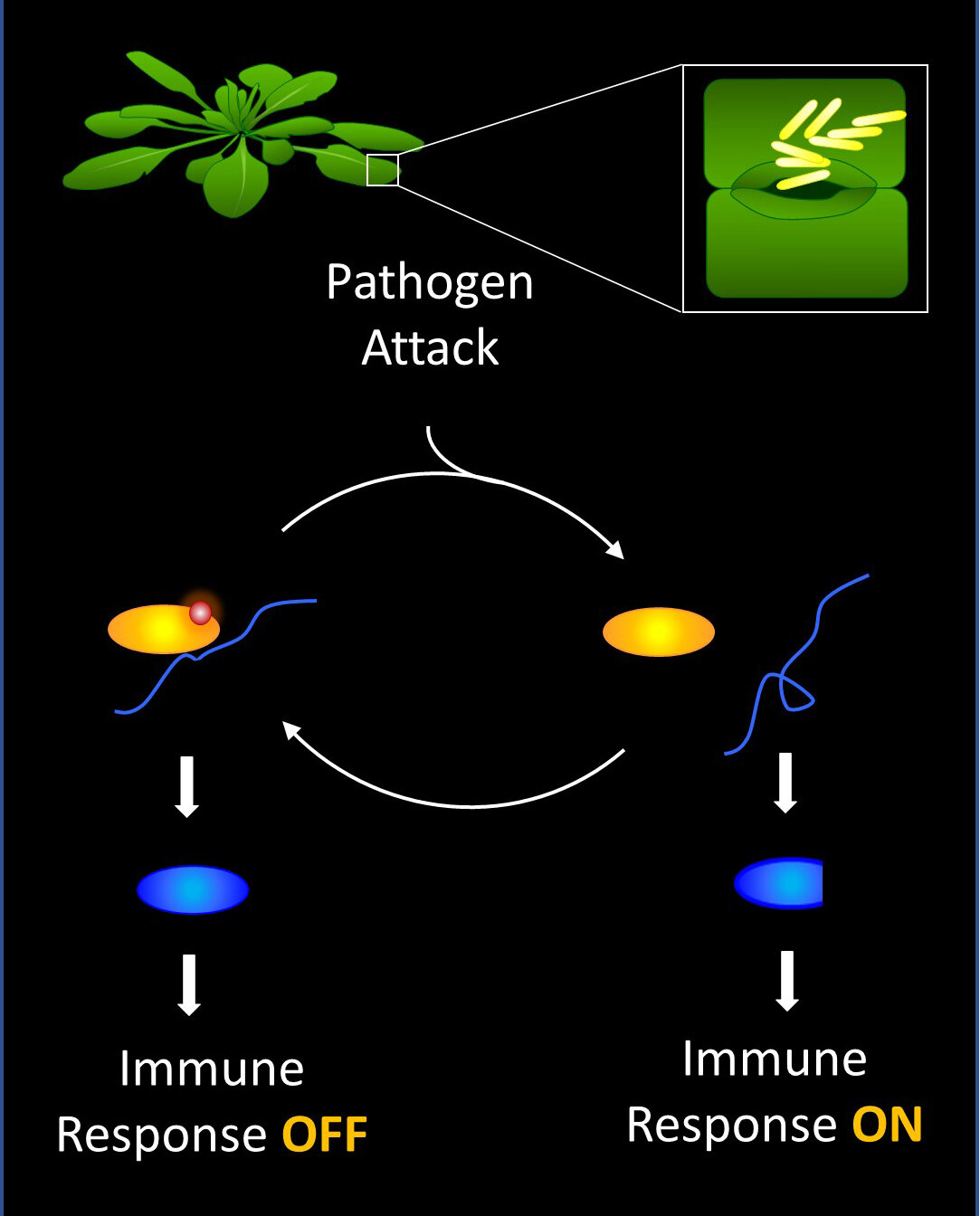
Till now, it’s very evident that almost every person must be knowing about the lethal impact of COVID-19 across the globe. But hardly, anyone knows the reason that how COVID-19, which was once living harmlessly in wildlife crossed the species barrier, giving it an evolutionary edge.
Recently, scientists at Duke University have found out that there have been numerous silent mutations in the genetic code of the coronavirus which helped it get the evolutionary edge and made it thrive beyond wildlife, leading to a global pandemic. These silent mutations guided the folding of RNA molecules in a unique way when present in human cells, setting the stage for the global crisis. The study has been published in the journal Peer. J.
The study involved various statistical methods that help in identifying various adaptations that the virus underwent. Researchers analyzed the genome sequence of the SARS-CoV-2 virus and other related coronaviruses often found in bats and pangolins to find out the adaptive changes in SARS-CoV2. “We’re trying to figure out what made this virus so unique,” said Alejandro Berrio, lead author of the paper and researcher at Duke University.
The earlier research highlighted the presence of positive selection within a gene that is responsible for encoding “Spike protein” on the surface of SARS-CoV2 surface. This increases the ability of the virus to infect the new cells in humans more outrageously. The study also indicated the presence of certain mutations in the viral genome that must have altered the spike protein, which made it thrive more easily among humans leading to a global pandemic. But this is not enough, the researchers also found out various other aspects which were not studies in previous researches, highlighting the reason that COVID has become so lethal and infectious. One of the critical reasons was a silent mutation in the two very important regions of the SARS-CoV2 genome, also referred to as Nsp4 and Nsp16. These mutations have been considered to give COVID-19 an evolutionary edge when compared with other similar strains, without impacting the proteins they naturally encode.
The study helped in understanding that the silent mutation instead of affecting the resultant protein affected how the RNA folds up in 3D shape, which eventually affects its functions in human cells. The deeper insights about how exactly these mutations govern the changes I RNA structure is yet to be elucidated in more detail. But never the less, the present study has very well contributed to understanding the viral leap from wildlife to humans.
“Nsp4 and Nsp16 are among the first RNA molecules that are produced when the virus infects a new person,” Berrio said. “The spike protein doesn’t get expressed until later. So they could make a better therapeutic target because they appear earlier in the viral life cycle.”
“Viruses are constantly mutating and evolving,” Berrio said. “So it’s possible that a new strain of coronavirus capable of infecting other animals may come along that also has the potential to spread to people as SARS-CoV-2 did. We’ll need to be able to recognize it and make efforts to contain it early.”